Alu Lab
The purpose of this lab was to take samples of our cheek cells and use them to create a PCR (polymerase chain reaction) so we could identify if our DNA was Heterozygous, Homozygous Positive, or Homozygous Negative.
This lab was also used to find out how the alu gene is used. As it turns out, it's great in identifying genetics and ethnicities in human populations because certain races have the same alu gene length/type. It can be used to show how the human race spread across the earth (because of this scientists have found out civilization started around Africa, then india, and split into Europe and Asia).
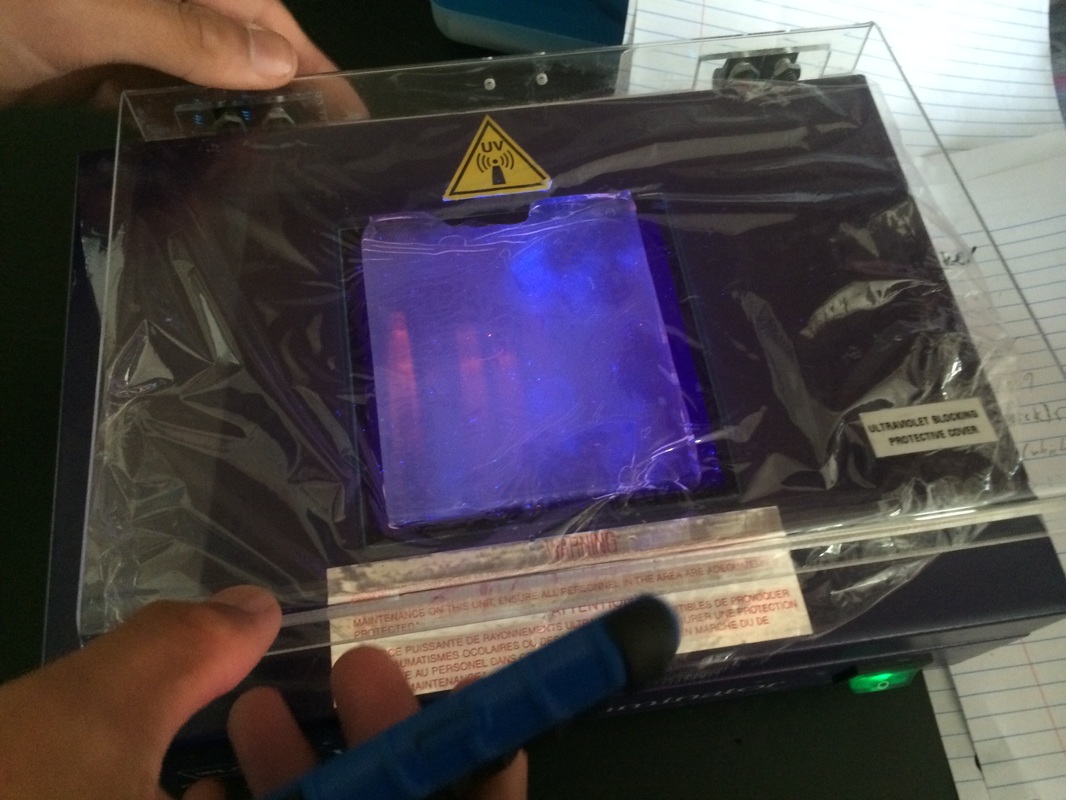
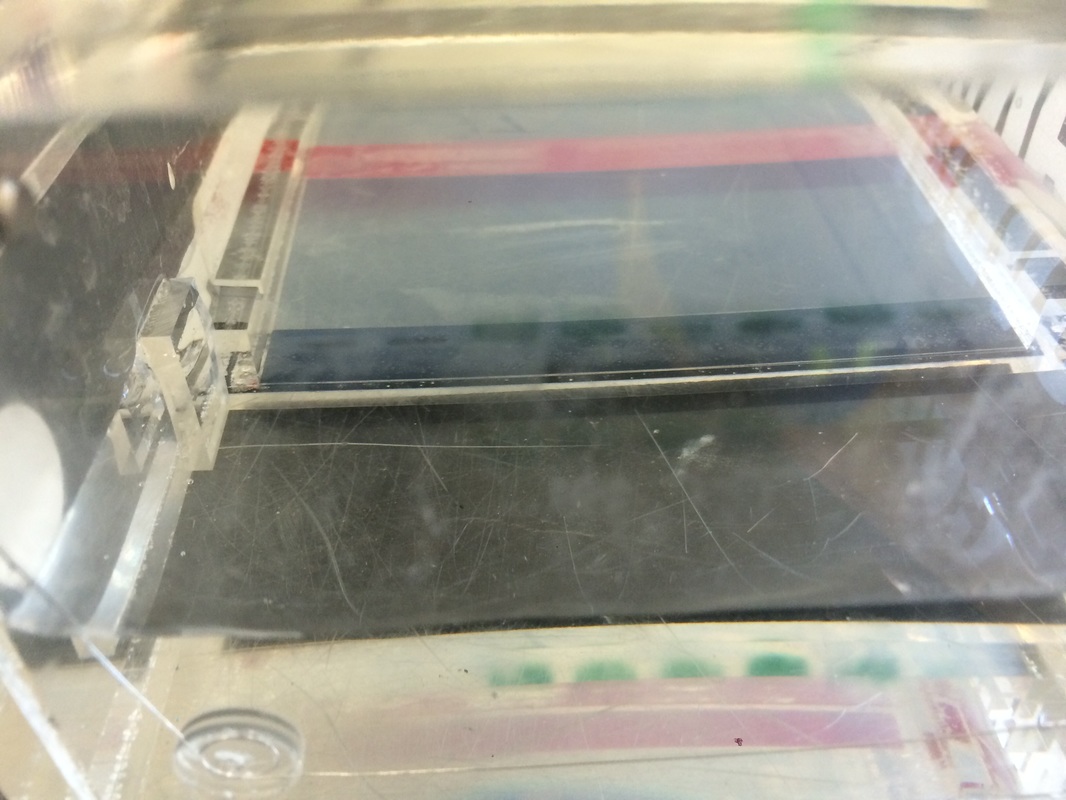
If we take cheek cells, remove metal ions, then break open the cells, we can force the DNA to create a PCR reaction and can view it with ultraviolet light. There is a tutorial showing the steps of PCR at http://www.dnalc.org/ddnalc/resources/pcr.html but eventually the DNA will either show up with a line at the top and bottom (heterozygous), just at top (homozygous positive), or just at bottom (homozygous negative). If no lines appear then human error most likely interfered. Being new to something like this, I can't really hypothesize what type I am unless I just guess without any logic. But right now I'm going to predict I'm heterozygous.
We needed many types of micropipets, a saline solution, Chelex, Master Mix, Primer Mix, a microcentrifuge, a thermal cycler, agarose gel, and ice or a refrigerator.
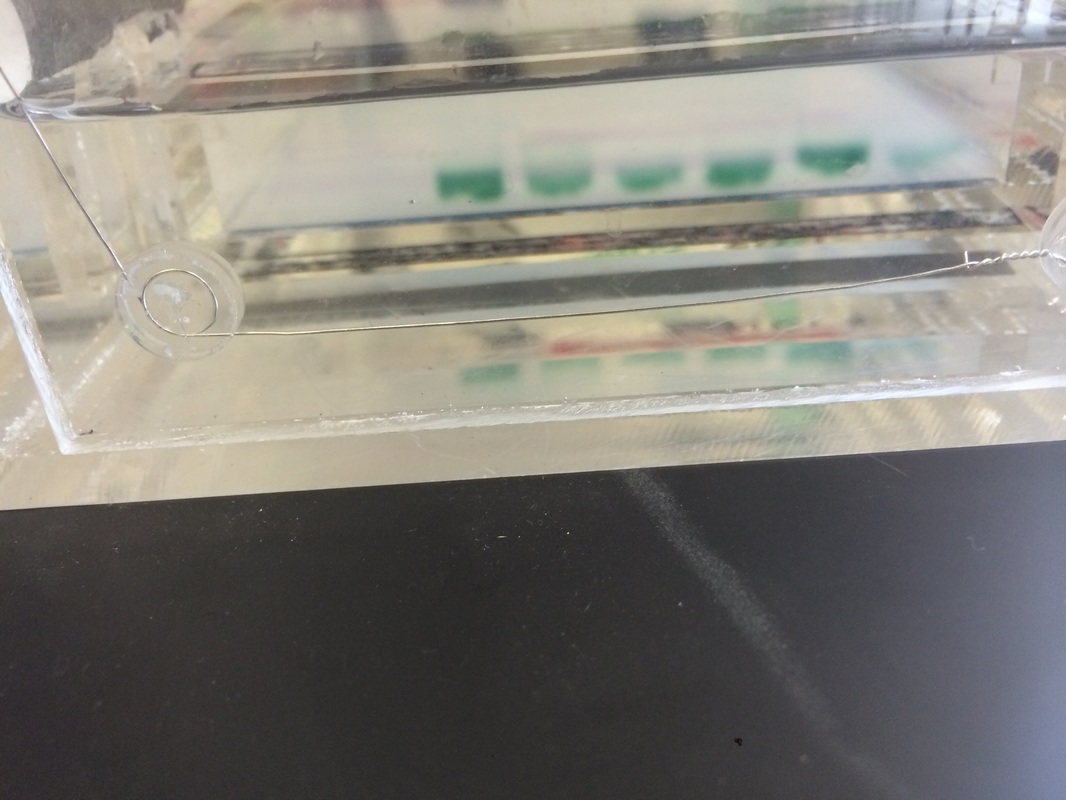
First we poured saline solution into our mouth and we swished it for about 30 seconds. We spit out the saline solution and using a micropipet, we transferred a small amount to a macrophage tube.Then we used a micro centrifuge and spun the cell suspension for a minute in order to pellet the cells. After getting rid of most of the supernatant, we then flicked or racked the pellet in order to "resuspend" the cell into an evenly mixed solution. Next, Chelex was added into the resuspended cell pellet and placed it on a thermal cycler for 10 minutes and placed it on the micro centrifuge for another minute. Now there was a bunch of Chelex beads and cell debris at the bottom. We had to be careful and micropipet the supernatant and NOT the debris. We proceeded to refrigerate the DNA until we combined 20 micrometers of Master Mix with 20 micrometers of Primer Mix, and then we added 10 micrometers of our DNA. We then made a gel and poked holes in it using a comb-like object. We inserted our DNA with great care and then used gel electrophoresis to analyze the results.
Unfortunately, while we were making the gel, someone accidentally let the gel leak out, so we had to borrow excess from some other group. This or some other form of human error caused our DNA to not show up that clearly when we tried to analyze it. Mine was one of the ones that weren't clear and I have no idea what I am to this day.
It's very likely that my DNA did not show up because I did something stupid and human error affected this, perhaps having too much supernatant, accidentally adding Chelex beads to the solution, or running out of Master/Primer Mix. Of course, it's very likely our spill affected this too, and I still believe that's that's the case.
Because human error affected the experiment, I cannot say if my hypothesis was right or not. To prevent errors like I explained above, we could have made extra sure about our measurements when we micropipetted. We also needed to make sure that the gel would not leak out of its container. Maybe if we made a second DNA tube as well, we could be more sure about our results.